![]()
![]() - ⇑
- ⇑

You need Javascript for this menu to work properly!
I've studied evolution in DNA and proteins, and malaria.

You need Javascript for this menu to work properly!

My goal is to develop statistical methods, probabilistic models, and algorithms for problems in computational biology. I am particularly interested in human disease and evolution, including the study of population genetics and protein sequences. I analyze large datasets and tackle problems with a variety of tools from statistics and computer science, balancing accuracy with efficiency. Examples include properly accounting for genomic relatedness to better associate genotype to disease, improving remote homology prediction across large protein databases, modeling psychiatric scores in clinical trials, and expanding functional annotations in malaria parasites.
I am an Assistant Professor in the Department of Biostatistics and Bioinformatics at Duke University. My research group, the Ochoa Lab, is part of the Center for Statistical Genetics and Genomics at Duke.
I was a Postdoctoral Reseach Associate in John D Storey's lab at Princeton University until 2018. I received my PhD in Molecular Biology in 2013 from Princeton University, coadvised by Mona Singh and Manuel Llinás. In 2006 I earned a double-majored in Biology and Mathematics at the Massachusetts Institute of Technology. See my CV (PDF).
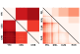

2021-01-19. Alejandro Ochoa, John D Storey. Estimating FST and kinship for arbitrary population structures. PLoS Genet. 17(1) e1009241. PubMed. PubMed Central. Article. bioRxiv 2016-10-27.
Alejandro Ochoa, John D Storey. FST and kinship for arbitrary population structures I: Generalized definitions. bioRxiv 2016-10-27.
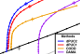

2017-04-12. Alejandro Ochoa, Mona Singh. Domain prediction with probabilistic directional context. Bioinf. 33(16) 2471-8. PubMed. PubMed Central. Article. bioRxiv 2016-12-14.
Software: dPUC2

2015-11-17. Alejandro Ochoa, John D Storey, Manuel Llinás, Mona Singh. Beyond the E-value: stratified statistics for protein domain prediction. PLoS Comput Biol. 11 e1004509. PubMed. PubMed Central. Article. arXiv 2014-09-23.
Software: DomStratStats and RandProt
2011-03-31. Alejandro Ochoa, Manuel Llinás, Mona Singh. Using context to improve protein domain identification. BMC Bioinformatics. 12:90. PubMed. PubMed Central. Article.
Software: dPUC1

2016-01-27. Simon A Cobbold, Joana M Santos, Alejandro Ochoa, David H Perlman, Manuel Llinás. Proteome-wide analysis reveals widespread lysine acetylation of major protein complexes in the malaria parasite. Sci Rep. 2016;6:19722. PubMed. PubMed Central. Article.
.

2013-11-13. Moriah L Szpara, Derek Gatherer, Alejandro Ochoa, Benjamin Greenbaum, Aidan Dolan, Rory J Bowden, Lynn W Enquist, Matthieu Legendre, Andrew J Davison. Evolution and diversity in human herpes simplex virus genomes. J Virol. 88:1209-27. PubMed. PubMed Central. Article.

2007-11-01. Gevorg Grigoryan, Alejandro Ochoa, Amy E Keating. Computing van der Waals energies in the context of the rotamer approximation. Proteins. 68(4) 863-78. PubMed. Article.
.